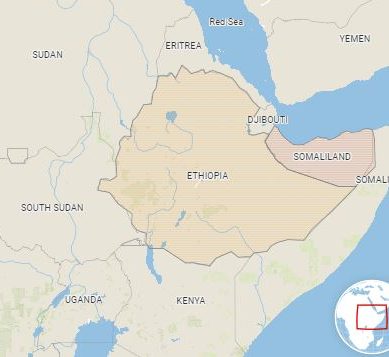
Mya Breitbart has hunted novel viruses in African termite mounds, Antarctic seals and water from the Red Sea. But to hit pay dirt, she has only to step into her back garden in Florida. Hanging around her swimming pool are spiny-backed orbweavers (Gasteracantha cancriformis) – striking spiders with bulbous white bodies, black speckles and six scarlet spikes that make them look like a piece of medieval weaponry.
Even more striking for Breitbart, a viral ecologist at the University of South Florida in St Petersburg, was what was inside. When she and her colleagues collected a few spiders and ground them up, they found two viruses previously unknown to science.
Although we humans have been focused on one particularly nasty virus since early 2020, there are legions of other viruses out there waiting to be discovered. Scientists estimate that there are about 10 trillion individual viral particles inhabiting the oceans alone at any given time – 10 billion times the estimated number of stars in the known Universe.
It’s becoming clear that ecosystems and organisms rely on viruses. Tiny but mighty, they have fuelled evolution for millions of years by shuttling genes between hosts. In the oceans, they slice open microorganisms, spilling their contents into the sea and flooding the food web with nutrients.
“Without viruses,” says Curtis Suttle, a virologist at the University of British Columbia in Vancouver, Canada, “we would not be alive.”
There are just 9,110 named species listed by the International Committee on Taxonomy of Viruses (ICTV), but that’s obviously a pitiful fraction of the total. In part, that’s because officially classifying a virus used to require scientists to culture a virus in its host or host cells – a time-consuming if not impossible process.
It’s also because the search has been biased towards viruses that cause diseases in humans or organisms we care about, such as farm animals and crop plants. Yet, as the Covid-19 pandemic has reminded us, it’s important to understand viruses that might jump from one host to another, threatening us, our animals or our crops.
Over the past ten years, the number of known and named viruses has exploded, owing to advances in the technology for finding them, plus a recent change to the rules for identifying new species, to allow naming without having to culture virus and host.
One of the most influential techniques is metagenomics, which allows researchers to sample the genomes in an environment without having to culture individual viruses. Newer technologies, such as single-virus sequencing, are adding even more viruses to the list, including some that are surprisingly common yet remained hidden until now. It’s an exciting time to be doing this kind of research, says Breitbart. “I think, in many ways, now is the time of the virome.”
In 2020 alone, the ICTV added 1,044 species to its official list and thousands more await description and naming. This proliferation of genomes prompted virologists to rethink the way they classify viruses and helped to clarify their evolution. There is strong evidence that viruses emerged multiple times, rather than sprouting from a single origin.
Even so, the true range of the viral world remains mostly uncharted, says Jens Kuhn, a virologist at the US National Institute of Allergy and Infectious Diseases facility at Fort Detrick, Maryland. “We really have absolutely no idea what’s out there.”
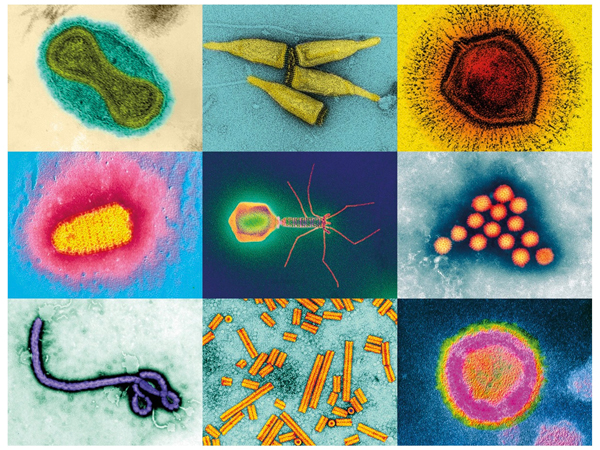
All viruses have two things in common: each encases its genome in a protein-based shell, and each relies on its host – be it a person, spider or plant – to reproduce itself. But beyond that general pattern lie endless variations.
There are minuscule circoviruses with only two or three genes, and massive mimiviruses that are bigger than some bacteria and carry hundreds of genes. There are lunar-lander-looking phage that infect bacteria and, of course, the killer spiky balls the world is now painfully familiar with.
There are viruses that store their genes as DNA and others that use RNA; there’s even a phage that uses an alternative genetic alphabet, replacing the chemical base A in the standard ACGT system with a different molecule, designated Z.
Viruses are so ubiquitous that they can turn up even when scientists aren’t looking for them. Frederik Schulz did not intend to study viruses as he pored over genome sequences from waste water. As a graduate student at the University of Vienna, in 2015 he was using metagenomics to hunt for bacteria. This involves isolating DNA from a whole mix of organisms, chopping it into bits and sequencing all of them.
A computer program then assembles the bits into individual genomes; it’s like solving hundreds of jigsaw puzzles whose pieces have been jumbled up.
Among the bacterial genomes, Schulz couldn’t help but notice a whopper of a virus genome – obvious because it carried genes for a viral shell – with a remarkable 1.57 million base pairs. It turned out to be a giant virus, part of a group whose members are large in terms of both genome size and absolute size (typically, 200 nanometres or more across).
These viruses infect amoebae, algae and other protists, putting them in a position to influence ecosystems both aquatic and terrestrial.
Schulz, now a microbiologist at the US Department of Energy Joint Genome Institute in Berkeley, California, decided to search for related viruses in metagenome data sets.
In 2020, in a single paper, he and his colleagues described more than 2,000 genomes from the group that contains giant viruses; before that, just 205 such genomes had been deposited in public databases.
Virologists have also looked inwards to find new species. Viral bioinformatician Luis Camarillo-Guerrero worked with colleagues at the Wellcome Sanger Institute in Hinxton, UK, to analyse metagenomes from the human gut, and built a database containing more than 140,000 kinds of phage.
More than half of these were new to science. Their study, published in February, matched others’ findings that one of the most common viruses to infect the bacteria in our guts is a group known as crAssphage (named after the cross-assembly software that picked it up in 2014).
Despite its abundance, not much is known about how it contributes to our microbiome, says Camarillo-Guerrero, who now works at DNA-sequencing company Illumina in Cambridge, UK.
Metagenomics has turned up a wealth of viruses, but it ignores many, too. RNA viruses aren’t sequenced in typical metagenomes, so microbiologist Colin Hill at University College Cork, Ireland, and his colleagues looked for them in databases of RNAs, called metatranscriptomes.
Scientists normally use these data to understand the genes in a population that are actively being turned into messenger RNA in to make proteins, but RNA virus genomes can show up, too. Using computational techniques to pull sequences out of the data, the team found 1,015 viral genomes in metatrancriptomes from sludge and water samples. Again, they’d massively increased the number of known viruses with a single paper.
Although it’s possible for these techniques to accidentally assemble genomes that aren’t real, researchers have quality-control techniques to guard against this. But there are other blind spots. For instance, viral species whose members are very diverse are fiendishly difficult to find because it’s hard for computer programs to piece together the disparate sequences.
The alternative is to sequence viral genomes one at a time, as microbiologist Manuel Martinez-Garcia does at the University of Alicante, Spain. He decided to try trickling seawater through a sorting machine to isolate single viruses, amplified their DNA, and got down to sequencing.
On his first attempt, he found 44 genomes. One turned out to represent some of the most abundant viruses in the ocean. This virus is so diverse – its genetic jigsaw pieces so varied from one virus particle to the next – that its genome had never popped up in metagenomics studies.
The team calls it 37-F6, for its location on the original laboratory dish, but Martinez-Garcia jokes that, given its ability to hide in plain sight, it should have been named 007, after fictional superspy James Bond.
- A Nature magazine report